Our research
Protein aggregation diseases, including Alzheimer’s Disease and Type 2 Diabetes, affect more than 50 million people in the US alone. However, the process by which proteins aggregate, and cells regulate that aggregation is still unclear. The goal of our research is to understand the large-scale protein conformational changes that proteins undergo during aggregation—a protein’s aggregation trajectory— and how cells control those trajectories. We are interested in understanding protein aggregation processes from ensemble biases in monomeric proteins to determinants of fibril polymorphism. We think that a biophysical/energy landscape understanding of protein aggregation is essential for understanding both the causes of protein aggregation diseases and for unraveling the non-Anfinsen protein behavior that is emblematic of aggregation-prone proteins including tau, alpha-synuclein, and others.
Our goal is to design research projects that change the way we think about those problems, and provide a unique lens to long-standing research questions. We value collaboration, innovation, and working from first principles.
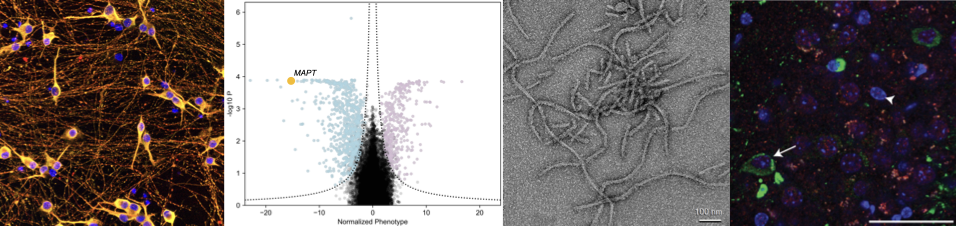
Cell-type specific regulation of the ubiquitin/proteasome system (UPS)
We are interested in understanding how the UPS of cells vulnerable and resilient to protein aggregation differ. We have identified new principles of tau processing by the UPS, including a new tau E3 ubiquitin ligase, CRL5SOCS4, and a role for oxidative stress in generating proteasome-derived tau proteolytic fragments. We use CRISPR-based genetic approaches combined with biochemical and biophysical techniques to reveal these mechanisms.
Sequence and cellular regulation of aggregation-prone proteins
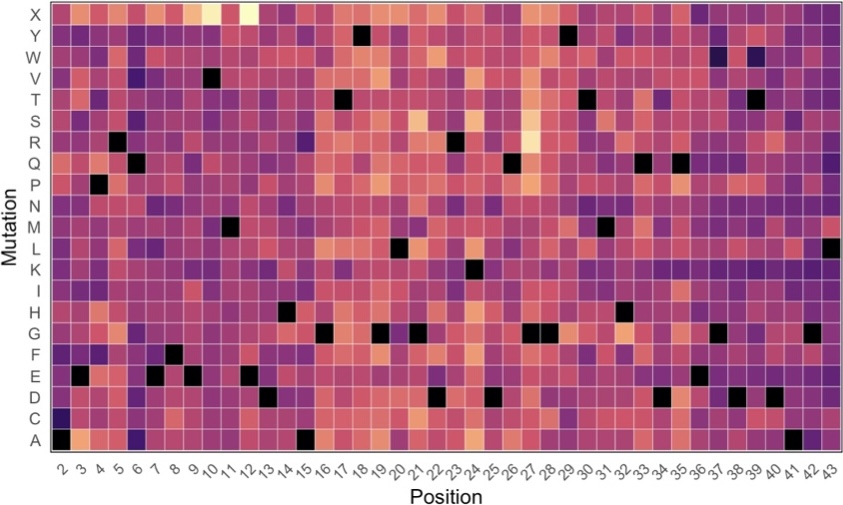 The most common proteins that aggregate in disease, like tau, are intrinsically disordered proteins (IDPs). IDPs do not populate a single three-dimensional structure like most proteins, but instead populate an ensemble: a collection of rapidly interconverting heterogeneous conformations. Small changes in these intrinsic (sequence-encoded) and extrinsic (cellular environment) factors can result in large changes to IDP ensembles. We are interested in understanding how sequence-encoded factors as well as cellular environmental factors alter IDP ensembles and aggregation trajectories. We take interdisciplinary collaborative approaches in our studied proteins’ native environment that combine high-resolution structural and biophysical techniques, high-throughput structural biology, and biochemistry and cell biology to gain a conformation-to-cell-state picture of IDP behavior.
The most common proteins that aggregate in disease, like tau, are intrinsically disordered proteins (IDPs). IDPs do not populate a single three-dimensional structure like most proteins, but instead populate an ensemble: a collection of rapidly interconverting heterogeneous conformations. Small changes in these intrinsic (sequence-encoded) and extrinsic (cellular environment) factors can result in large changes to IDP ensembles. We are interested in understanding how sequence-encoded factors as well as cellular environmental factors alter IDP ensembles and aggregation trajectories. We take interdisciplinary collaborative approaches in our studied proteins’ native environment that combine high-resolution structural and biophysical techniques, high-throughput structural biology, and biochemistry and cell biology to gain a conformation-to-cell-state picture of IDP behavior.
New models of neurodegeneration and aging
Current models of neurodegeneration and aging have to balance speed, molecular resoltuion, scalability, and recapitulation of disease and aging phenotypes. What if we did not have to choose? Our lab is building new tools for iPSC systems, new models of disease-vulnerable cell types, and CRISPR-based tools for recapitulting key transcriptional and phenotypic signatures of aging and neurodegeneration.